Chapter 1 R and the Tidyverse
1.1 Overview
This chapter provides an introduction to data science and the R programming language. The goal here is to get your hands dirty right from the start! We will walk through an entire data analysis, and along the way introduce different types of data analysis question, some fundamental programming concepts in R, and the basics of loading, cleaning, and visualizing data. In the following chapters, we will dig into each of these steps in much more detail; but for now, let’s jump in to see how much we can do with data science!
1.2 Chapter learning objectives
By the end of the chapter, readers will be able to do the following:
- Identify the different types of data analysis question and categorize a question into the correct type.
- Load the
tidyversepackage into R. - Read tabular data with
read_csv. - Create new variables and objects in R using the assignment symbol.
- Create and organize subsets of tabular data using
filter,select,arrange, andslice. - Add and modify columns in tabular data using
mutate. - Visualize data with a
ggplotbar plot. - Use
?to access help and documentation tools in R.
1.3 Canadian languages data set
In this chapter, we will walk through a full analysis of a data set relating to languages spoken at home by Canadian residents (Figure 1.1). Many Indigenous peoples exist in Canada with their own cultures and languages; these languages are often unique to Canada and not spoken anywhere else in the world (Statistics Canada 2018). Sadly, colonization has led to the loss of many of these languages. For instance, generations of children were not allowed to speak their mother tongue (the first language an individual learns in childhood) in Canadian residential schools. Colonizers also renamed places they had “discovered” (K. Wilson 2018). Acts such as these have significantly harmed the continuity of Indigenous languages in Canada, and some languages are considered “endangered” as few people report speaking them. To learn more, please see Canadian Geographic’s article, “Mapping Indigenous Languages in Canada” (Walker 2017), They Came for the Children: Canada, Aboriginal peoples, and Residential Schools (Truth and Reconciliation Commission of Canada 2012) and the Truth and Reconciliation Commission of Canada’s Calls to Action (Truth and Reconciliation Commission of Canada 2015).
Figure 1.1: Map of Canada.
The data set we will study in this chapter is taken from
the canlang R data package
(Timbers 2020), which has
population language data collected during the 2016 Canadian census (Statistics Canada 2016a).
In this data, there are 214 languages recorded, each having six different properties:
category: Higher-level language category, describing whether the language is an Official Canadian language, an Aboriginal (i.e., Indigenous) language, or a Non-Official and Non-Aboriginal language.language: The name of the language.mother_tongue: Number of Canadian residents who reported the language as their mother tongue. Mother tongue is generally defined as the language someone was exposed to since birth.most_at_home: Number of Canadian residents who reported the language as being spoken most often at home.most_at_work: Number of Canadian residents who reported the language as being used most often at work.lang_known: Number of Canadian residents who reported knowledge of the language.
According to the census, more than 60 Aboriginal languages were reported as being spoken in Canada. Suppose we want to know which are the most common; then we might ask the following question, which we wish to answer using our data:
Which ten Aboriginal languages were most often reported in 2016 as mother tongues in Canada, and how many people speak each of them?
Note: Data science cannot be done without a deep understanding of the data and problem domain. In this book, we have simplified the data sets used in our examples to concentrate on methods and fundamental concepts. But in real life, you cannot and should not do data science without a domain expert. Alternatively, it is common to practice data science in your own domain of expertise! Remember that when you work with data, it is essential to think about how the data were collected, which affects the conclusions you can draw. If your data are biased, then your results will be biased!
1.4 Asking a question
Every good data analysis begins with a question—like the above—that you aim to answer using data. As it turns out, there are actually a number of different types of question regarding data: descriptive, exploratory, predictive, inferential, causal, and mechanistic, all of which are defined in Table 1.1. Carefully formulating a question as early as possible in your analysis—and correctly identifying which type of question it is—will guide your overall approach to the analysis as well as the selection of appropriate tools.
| Question type | Description | Example |
|---|---|---|
| Descriptive | A question that asks about summarized characteristics of a data set without interpretation (i.e., report a fact). | How many people live in each province and territory in Canada? |
| Exploratory | A question that asks if there are patterns, trends, or relationships within a single data set. Often used to propose hypotheses for future study. | Does political party voting change with indicators of wealth in a set of data collected on 2,000 people living in Canada? |
| Predictive | A question that asks about predicting measurements or labels for individuals (people or things). The focus is on what things predict some outcome, but not what causes the outcome. | What political party will someone vote for in the next Canadian election? |
| Inferential | A question that looks for patterns, trends, or relationships in a single data set and also asks for quantification of how applicable these findings are to the wider population. | Does political party voting change with indicators of wealth for all people living in Canada? |
| Causal | A question that asks about whether changing one factor will lead to a change in another factor, on average, in the wider population. | Does wealth lead to voting for a certain political party in Canadian elections? |
| Mechanistic | A question that asks about the underlying mechanism of the observed patterns, trends, or relationships (i.e., how does it happen?) | How does wealth lead to voting for a certain political party in Canadian elections? |
In this book, you will learn techniques to answer the first four types of question: descriptive, exploratory, predictive, and inferential; causal and mechanistic questions are beyond the scope of this book. In particular, you will learn how to apply the following analysis tools:
- Summarization: computing and reporting aggregated values pertaining to a data set. Summarization is most often used to answer descriptive questions, and can occasionally help with answering exploratory questions. For example, you might use summarization to answer the following question: What is the average race time for runners in this data set? Tools for summarization are covered in detail in Chapters 2 and 3, but appear regularly throughout the text.
- Visualization: plotting data graphically. Visualization is typically used to answer descriptive and exploratory questions, but plays a critical supporting role in answering all of the types of question in Table 1.1. For example, you might use visualization to answer the following question: Is there any relationship between race time and age for runners in this data set? This is covered in detail in Chapter 4, but again appears regularly throughout the book.
- Classification: predicting a class or category for a new observation. Classification is used to answer predictive questions. For example, you might use classification to answer the following question: Given measurements of a tumor’s average cell area and perimeter, is the tumor benign or malignant? Classification is covered in Chapters 5 and 6.
- Regression: predicting a quantitative value for a new observation. Regression is also used to answer predictive questions. For example, you might use regression to answer the following question: What will be the race time for a 20-year-old runner who weighs 50kg? Regression is covered in Chapters 7 and 8.
- Clustering: finding previously unknown/unlabeled subgroups in a data set. Clustering is often used to answer exploratory questions. For example, you might use clustering to answer the following question: What products are commonly bought together on Amazon? Clustering is covered in Chapter 9.
- Estimation: taking measurements for a small number of items from a large group and making a good guess for the average or proportion for the large group. Estimation is used to answer inferential questions. For example, you might use estimation to answer the following question: Given a survey of cellphone ownership of 100 Canadians, what proportion of the entire Canadian population own Android phones? Estimation is covered in Chapter 10.
Referring to Table 1.1, our question about Aboriginal languages is an example of a descriptive question: we are summarizing the characteristics of a data set without further interpretation. And referring to the list above, it looks like we should use visualization and perhaps some summarization to answer the question. So in the remainder of this chapter, we will work towards making a visualization that shows us the ten most common Aboriginal languages in Canada and their associated counts, according to the 2016 census.
1.5 Loading a tabular data set
A data set is, at its core essence, a structured collection of numbers and characters. Aside from that, there are really no strict rules; data sets can come in many different forms! Perhaps the most common form of data set that you will find in the wild, however, is tabular data. Think spreadsheets in Microsoft Excel: tabular data are rectangular-shaped and spreadsheet-like, as shown in Figure 1.2. In this book, we will focus primarily on tabular data.
Since we are using R for data analysis in this book, the first step for us is to load the data into R. When we load tabular data into R, it is represented as a data frame object. Figure 1.2 shows that an R data frame is very similar to a spreadsheet. We refer to the rows as observations; these are the individual objects for which we collect data. In Figure 1.2, the observations are languages. We refer to the columns as variables; these are the characteristics of each observation. In Figure 1.2, the variables are the the language’s category, its name, the number of mother tongue speakers, etc.
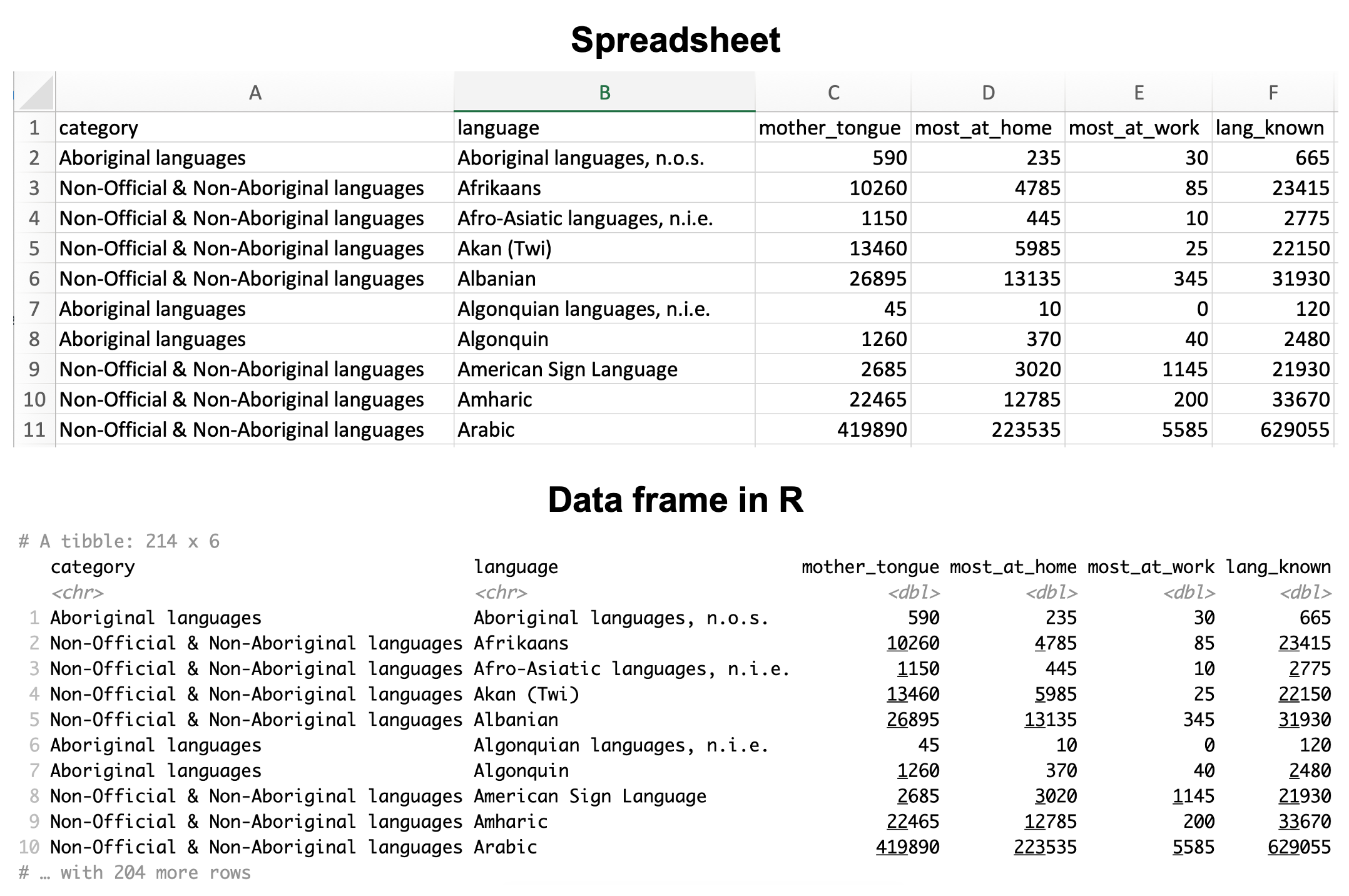
Figure 1.2: A spreadsheet versus a data frame in R.
The first kind of data file that we will learn how to load into R as a data
frame is the comma-separated values format (.csv for short). These files
have names ending in .csv, and can be opened and saved using common
spreadsheet programs like Microsoft Excel and Google Sheets. For example, the
.csv file named can_lang.csv
is included with the code for this book.
If we were to open this data in a plain text editor (a program like Notepad that just shows
text with no formatting), we would see each row on its own line, and each entry in the table separated by a comma:
category,language,mother_tongue,most_at_home,most_at_work,lang_known
Aboriginal languages,"Aboriginal languages, n.o.s.",590,235,30,665
Non-Official & Non-Aboriginal languages,Afrikaans,10260,4785,85,23415
Non-Official & Non-Aboriginal languages,"Afro-Asiatic languages, n.i.e.",1150,44
Non-Official & Non-Aboriginal languages,Akan (Twi),13460,5985,25,22150
Non-Official & Non-Aboriginal languages,Albanian,26895,13135,345,31930
Aboriginal languages,"Algonquian languages, n.i.e.",45,10,0,120
Aboriginal languages,Algonquin,1260,370,40,2480
Non-Official & Non-Aboriginal languages,American Sign Language,2685,3020,1145,21
Non-Official & Non-Aboriginal languages,Amharic,22465,12785,200,33670To load this data into R so that we can do things with it (e.g., perform
analyses or create data visualizations), we will need to use a function. A
function is a special word in R that takes instructions (we call these
arguments) and does something. The function we will use to load a .csv file
into R is called read_csv. In its most basic
use-case, read_csv expects that the data file:
- has column names (or headers),
- uses a comma (
,) to separate the columns, and - does not have row names.
Below you’ll see the code used to load the data into R using the read_csv
function. Note that the read_csv function is not included in the base
installation of R, meaning that it is not one of the primary functions ready to
use when you install R. Therefore, you need to load it from somewhere else
before you can use it. The place from which we will load it is called an R package.
An R package is a collection of functions that can be used in addition to the
built-in R package functions once loaded. The read_csv function, in
particular, can be made accessible by loading
the tidyverse R package (Wickham 2021b; Wickham et al. 2019)
using the library function. The tidyverse package contains many
functions that we will use throughout this book to load, clean, wrangle,
and visualize data.
## ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
## ✔ dplyr 1.1.2 ✔ readr 2.1.4
## ✔ forcats 1.0.0 ✔ stringr 1.5.0
## ✔ ggplot2 3.4.2 ✔ tibble 3.2.1
## ✔ lubridate 1.9.2 ✔ tidyr 1.3.0
## ✔ purrr 1.0.1
## ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
## ✖ dplyr::filter() masks stats::filter()
## ✖ dplyr::lag() masks stats::lag()
## ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorsNote: You may have noticed that we got some extra output from R regarding attached packages and conflicts below our code line. These are examples of messages in R, which give the user more information that might be handy to know. The
Attaching packagesmessage is natural when loadingtidyverse, sincetidyverseactually automatically causes other packages to be imported too, such asdplyr. In the future, when we loadtidyversein this book, we will silence these messages to help with the readability of the book. TheConflictsmessage is also totally normal in this circumstance. This message tells you if functions from different packages share the same name, which is confusing to R. For example, in this case, thedplyrpackage and thestatspackage both provide a function calledfilter. The message above (dplyr::filter() masks stats::filter()) is R telling you that it is going to default to thedplyrpackage version of this function. So if you use thefilterfunction, you will be using thedplyrversion. In order to use thestatsversion, you need to use its full namestats::filter. Messages are not errors, so generally you don’t need to take action when you see a message; but you should always read the message and critically think about what it means and whether you need to do anything about it.
After loading the tidyverse package, we can call the read_csv function and
pass it a single argument: the name of the file, "can_lang.csv". We have to
put quotes around file names and other letters and words that we use in our
code to distinguish it from the special words (like functions!) that make up the R programming
language. The file’s name is the only argument we need to provide because our
file satisfies everything else that the read_csv function expects in the default
use-case. Figure 1.3 describes how we use the read_csv
to read data into R.

Figure 1.3: Syntax for the read_csv function.
## # A tibble: 214 × 6
## category language mother_tongue most_at_home most_at_work lang_known
## <chr> <chr> <dbl> <dbl> <dbl> <dbl>
## 1 Aboriginal langu… Aborigi… 590 235 30 665
## 2 Non-Official & N… Afrikaa… 10260 4785 85 23415
## 3 Non-Official & N… Afro-As… 1150 445 10 2775
## 4 Non-Official & N… Akan (T… 13460 5985 25 22150
## 5 Non-Official & N… Albanian 26895 13135 345 31930
## 6 Aboriginal langu… Algonqu… 45 10 0 120
## 7 Aboriginal langu… Algonqu… 1260 370 40 2480
## 8 Non-Official & N… America… 2685 3020 1145 21930
## 9 Non-Official & N… Amharic 22465 12785 200 33670
## 10 Non-Official & N… Arabic 419890 223535 5585 629055
## # ℹ 204 more rowsNote: There is another function that also loads csv files named
read.csv. We will always useread_csvin this book, as it is designed to play nicely with all of the othertidyversefunctions, which we will use extensively. Be careful not to accidentally useread.csv, as it can cause some tricky errors to occur in your code that are hard to track down!
1.6 Naming things in R
When we loaded the 2016 Canadian census language data
using read_csv, we did not give this data frame a name.
Therefore the data was just printed on the screen,
and we cannot do anything else with it. That isn’t very useful.
What would be more useful would be to give a name
to the data frame that read_csv outputs,
so that we can refer to it later for analysis and visualization.
The way to assign a name to a value in R is via the assignment symbol <-.
On the left side of the assignment symbol you put the name that you want
to use, and on the right side of the assignment symbol
you put the value that you want the name to refer to.
Names can be used to refer to almost anything in R, such as numbers,
words (also known as strings of characters), and data frames!
Below, we set my_number to 3 (the result of 1+2)
and we set name to the string "Alice".
Note that when
we name something in R using the assignment symbol, <-,
we do not need to surround the name we are creating with quotes. This is
because we are formally telling R that this special word denotes
the value of whatever is on the right-hand side.
Only characters and words that act as values on the right-hand side of the assignment
symbol—e.g., the file name "data/can_lang.csv" that we specified before, or "Alice" above—need
to be surrounded by quotes.
After making the assignment, we can use the special name words we have created in
place of their values. For example, if we want to do something with the value 3 later on,
we can just use my_number instead. Let’s try adding 2 to my_number; you will see that
R just interprets this as adding 3 and 2:
## [1] 5Object names can consist of letters, numbers, periods . and underscores _.
Other symbols won’t work since they have their own meanings in R. For example,
- is the subtraction symbol; if we try to assign a name with
the - symbol, R will complain and we will get an error!
Error in my - number <- 1 : object 'my' not foundThere are certain conventions for naming objects in R.
When naming an object we
suggest using only lowercase letters, numbers and underscores _ to separate
the words in a name. R is case sensitive, which means that Letter and
letter would be two different objects in R. You should also try to give your
objects meaningful names. For instance, you can name a data frame x.
However, using more meaningful terms, such as language_data, will help you
remember what each name in your code represents. We recommend following the
Tidyverse naming conventions outlined in the Tidyverse Style Guide (Wickham 2020). Let’s
now use the assignment symbol to give the name
can_lang to the 2016 Canadian census language data frame that we get from
read_csv.
Wait a minute, nothing happened this time! Where’s our data?
Actually, something did happen: the data was loaded in
and now has the name can_lang associated with it.
And we can use that name to access the data frame and do things with it.
For example, we can type the name of the data frame to print the first few rows
on the screen. You will also see at the top that the number of observations (i.e., rows) and
variables (i.e., columns) are printed. Printing the first few rows of a data frame
like this is a handy way to get a quick sense for what is contained in a data frame.
## # A tibble: 214 × 6
## category language mother_tongue most_at_home most_at_work lang_known
## <chr> <chr> <dbl> <dbl> <dbl> <dbl>
## 1 Aboriginal langu… Aborigi… 590 235 30 665
## 2 Non-Official & N… Afrikaa… 10260 4785 85 23415
## 3 Non-Official & N… Afro-As… 1150 445 10 2775
## 4 Non-Official & N… Akan (T… 13460 5985 25 22150
## 5 Non-Official & N… Albanian 26895 13135 345 31930
## 6 Aboriginal langu… Algonqu… 45 10 0 120
## 7 Aboriginal langu… Algonqu… 1260 370 40 2480
## 8 Non-Official & N… America… 2685 3020 1145 21930
## 9 Non-Official & N… Amharic 22465 12785 200 33670
## 10 Non-Official & N… Arabic 419890 223535 5585 629055
## # ℹ 204 more rows1.7 Creating subsets of data frames with filter & select
Now that we’ve loaded our data into R, we can start wrangling the data to
find the ten Aboriginal languages that were most often reported
in 2016 as mother tongues in Canada. In particular, we will construct
a table with the ten Aboriginal languages that have the largest
counts in the mother_tongue column.
The filter and select functions from the tidyverse package will help us
here. The filter function allows you to obtain a subset of the
rows with specific values, while the select function allows you
to obtain a subset of the columns. Therefore, we can filter the rows to extract the
Aboriginal languages in the data set, and then use select to obtain only the
columns we want to include in our table.
1.7.1 Using filter to extract rows
Looking at the can_lang data above, we see the category column contains different
high-level categories of languages, which include “Aboriginal languages”,
“Non-Official & Non-Aboriginal languages” and “Official languages”. To answer
our question we want to filter our data set so we restrict our attention
to only those languages in the “Aboriginal languages” category.
We can use the filter function to obtain the subset of rows with desired
values from a data frame. Figure 1.4 outlines what arguments we need to specify to use filter. The first argument to filter is the name of the data frame
object, can_lang. The second argument is a logical statement to use when
filtering the rows. A logical statement evaluates to either TRUE or FALSE;
filter keeps only those rows for which the logical statement evaluates to TRUE.
For example, in our analysis, we are interested in keeping only languages in the
“Aboriginal languages” higher-level category. We can use
the equivalency operator == to compare the values
of the category column with the value "Aboriginal languages"; you will learn about
many other kinds of logical statements in Chapter 3. Similar to
when we loaded the data file and put quotes around the file name, here we need
to put quotes around "Aboriginal languages". Using quotes tells R that this
is a string value and not one of the special words that make up the R
programming language, or one of the names we have given to data frames in the
code we have already written.
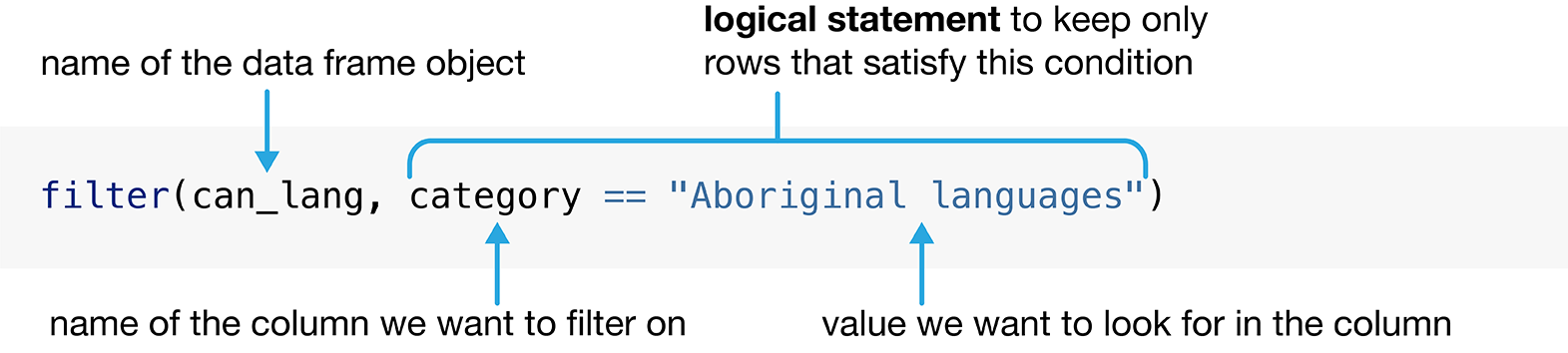
Figure 1.4: Syntax for the filter function.
With these arguments, filter returns a data frame that has all the columns of
the input data frame, but only those rows we asked for in our logical filter
statement.
## # A tibble: 67 × 6
## category language mother_tongue most_at_home most_at_work lang_known
## <chr> <chr> <dbl> <dbl> <dbl> <dbl>
## 1 Aboriginal langu… Aborigi… 590 235 30 665
## 2 Aboriginal langu… Algonqu… 45 10 0 120
## 3 Aboriginal langu… Algonqu… 1260 370 40 2480
## 4 Aboriginal langu… Athabas… 50 10 0 85
## 5 Aboriginal langu… Atikame… 6150 5465 1100 6645
## 6 Aboriginal langu… Babine … 110 20 10 210
## 7 Aboriginal langu… Beaver 190 50 0 340
## 8 Aboriginal langu… Blackfo… 2815 1110 85 5645
## 9 Aboriginal langu… Carrier 1025 250 15 2100
## 10 Aboriginal langu… Cayuga 45 10 10 125
## # ℹ 57 more rowsIt’s good practice to check the output after using a
function in R. We can see the original can_lang data set contained 214 rows
with multiple kinds of category. The data frame
aboriginal_lang contains only 67 rows, and looks like it only contains languages in
the “Aboriginal languages” in the category column. So it looks like the function
gave us the result we wanted!
1.7.2 Using select to extract columns
Now let’s use select to extract the language and mother_tongue columns
from this data frame. Figure 1.5 shows us the syntax for the select function. To extract these columns, we need to provide the select
function with three arguments. The first argument is the name of the data frame
object, which in this example is aboriginal_lang. The second and third
arguments are the column names that we want to select: language and
mother_tongue. After passing these three arguments, the select function
returns two columns (the language and mother_tongue columns that we asked
for) as a data frame. This code is also a great example of why being
able to name things in R is useful: you can see that we are using the
result of our earlier filter step (which we named aboriginal_lang) here
in the next step of the analysis!

Figure 1.5: Syntax for the select function.
## # A tibble: 67 × 2
## language mother_tongue
## <chr> <dbl>
## 1 Aboriginal languages, n.o.s. 590
## 2 Algonquian languages, n.i.e. 45
## 3 Algonquin 1260
## 4 Athabaskan languages, n.i.e. 50
## 5 Atikamekw 6150
## 6 Babine (Wetsuwet'en) 110
## 7 Beaver 190
## 8 Blackfoot 2815
## 9 Carrier 1025
## 10 Cayuga 45
## # ℹ 57 more rows1.8 Using arrange to order and slice to select rows by index number
We have used filter and select to obtain a table with only the Aboriginal
languages in the data set and their associated counts. However, we want to know
the ten languages that are spoken most often. As a next step, we will
order the mother_tongue column from largest to smallest value and then extract only
the top ten rows. This is where the arrange and slice functions come to the
rescue!
The arrange function allows us to order the rows of a data frame by the
values of a particular column. Figure 1.6 details what arguments we need to specify to
use the arrange function. We need to pass the data frame as the first
argument to this function, and the variable to order by as the second argument.
Since we want to choose the ten Aboriginal languages most often reported as a mother tongue
language, we will use the arrange function to order the rows in our
selected_lang data frame by the mother_tongue column. We want to
arrange the rows in descending order (from largest to smallest),
so we pass the column to the desc function before using it as an argument.
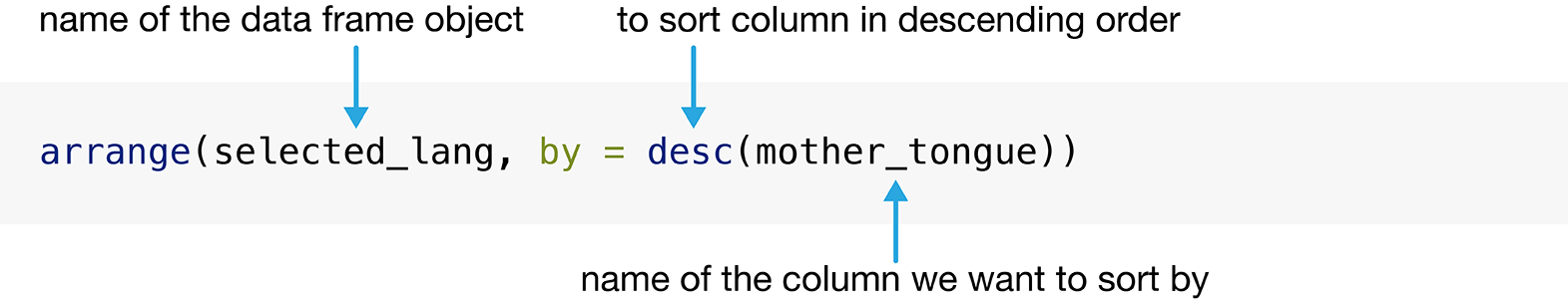

Figure 1.6: Syntax for the arrange function.
## # A tibble: 67 × 2
## language mother_tongue
## <chr> <dbl>
## 1 Cree, n.o.s. 64050
## 2 Inuktitut 35210
## 3 Ojibway 17885
## 4 Oji-Cree 12855
## 5 Dene 10700
## 6 Montagnais (Innu) 10235
## 7 Mi'kmaq 6690
## 8 Atikamekw 6150
## 9 Plains Cree 3065
## 10 Stoney 3025
## # ℹ 57 more rowsNext we will use the slice function, which selects rows according to their
row number. Since we want to choose the most common ten languages, we will indicate we want the
rows 1 to 10 using the argument 1:10.
## # A tibble: 10 × 2
## language mother_tongue
## <chr> <dbl>
## 1 Cree, n.o.s. 64050
## 2 Inuktitut 35210
## 3 Ojibway 17885
## 4 Oji-Cree 12855
## 5 Dene 10700
## 6 Montagnais (Innu) 10235
## 7 Mi'kmaq 6690
## 8 Atikamekw 6150
## 9 Plains Cree 3065
## 10 Stoney 30251.9 Adding and modifying columns using mutate
Recall that our data analysis question referred to the count of Canadians
that speak each of the top ten most commonly reported Aboriginal languages as
their mother tongue, and the ten_lang data frame indeed contains those
counts… But perhaps, seeing these numbers, we became curious about the
percentage of the population of Canada associated with each count. It is
common to come up with new data analysis questions in the process of answering
a first one—so fear not and explore! To answer this small
question along the way, we need to divide each count in the mother_tongue
column by the total Canadian population according to the 2016
census—i.e., 35,151,728—and multiply it by 100. We can perform
this computation using the mutate function. We pass the ten_lang
data frame as its first argument, then specify the equation that computes the percentages
in the second argument. By using a new variable name on the left hand side of the equation,
we will create a new column in the data frame; and if we use an existing name, we will
modify that variable. In this case, we will opt to
create a new column called mother_tongue_percent.
canadian_population = 35151728
ten_lang_percent = mutate(ten_lang, mother_tongue_percent = 100 * mother_tongue / canadian_population)
ten_lang_percent## # A tibble: 10 × 3
## language mother_tongue mother_tongue_percent
## <chr> <dbl> <dbl>
## 1 Cree, n.o.s. 64050 0.182
## 2 Inuktitut 35210 0.100
## 3 Ojibway 17885 0.0509
## 4 Oji-Cree 12855 0.0366
## 5 Dene 10700 0.0304
## 6 Montagnais (Innu) 10235 0.0291
## 7 Mi'kmaq 6690 0.0190
## 8 Atikamekw 6150 0.0175
## 9 Plains Cree 3065 0.00872
## 10 Stoney 3025 0.00861The ten_lang_percent data frame shows that
the ten Aboriginal languages in the ten_lang data frame were spoken
as a mother tongue by between 0.008% and 0.18% of the Canadian population.
1.10 Exploring data with visualizations
The ten_lang table we generated in Section 1.8 answers our initial data analysis question.
Are we done? Well, not quite; tables are almost never the best way to present
the result of your analysis to your audience. Even the ten_lang table with
only two columns presents some difficulty: for example, you have to scrutinize
the table quite closely to get a sense for the relative numbers of speakers of
each language. When you move on to more complicated analyses, this issue only
gets worse. In contrast, a visualization would convey this information in a much
more easily understood format.
Visualizations are a great tool for summarizing information to help you
effectively communicate with your audience, and
creating effective data visualizations is an essential component of any data
analysis. In this section we will develop a visualization of the
ten Aboriginal languages that were most often reported in 2016 as mother tongues in
Canada, as well as the number of people that speak each of them.
1.10.1 Using ggplot to create a bar plot
In our data set, we can see that language and mother_tongue are in separate
columns (or variables). In addition, there is a single row (or observation) for each language.
The data are, therefore, in what we call a tidy data format. Tidy data is a
fundamental concept and will be a significant focus in the remainder of this
book: many of the functions from tidyverse require tidy data, including the
ggplot function that we will use shortly for our visualization. We will
formally introduce tidy data in Chapter 3.
We will make a bar plot to visualize our data. A bar plot is a chart where the
lengths of the bars represent certain values, like counts or proportions. We
will make a bar plot using the mother_tongue and language columns from our
ten_lang data frame. To create a bar plot of these two variables using the
ggplot function, we must specify the data frame, which variables
to put on the x and y axes, and what kind of plot to create. The ggplot
function and its common usage is illustrated in Figure 1.7.
Figure 1.8 shows the resulting bar plot
generated by following the instructions in Figure 1.7.
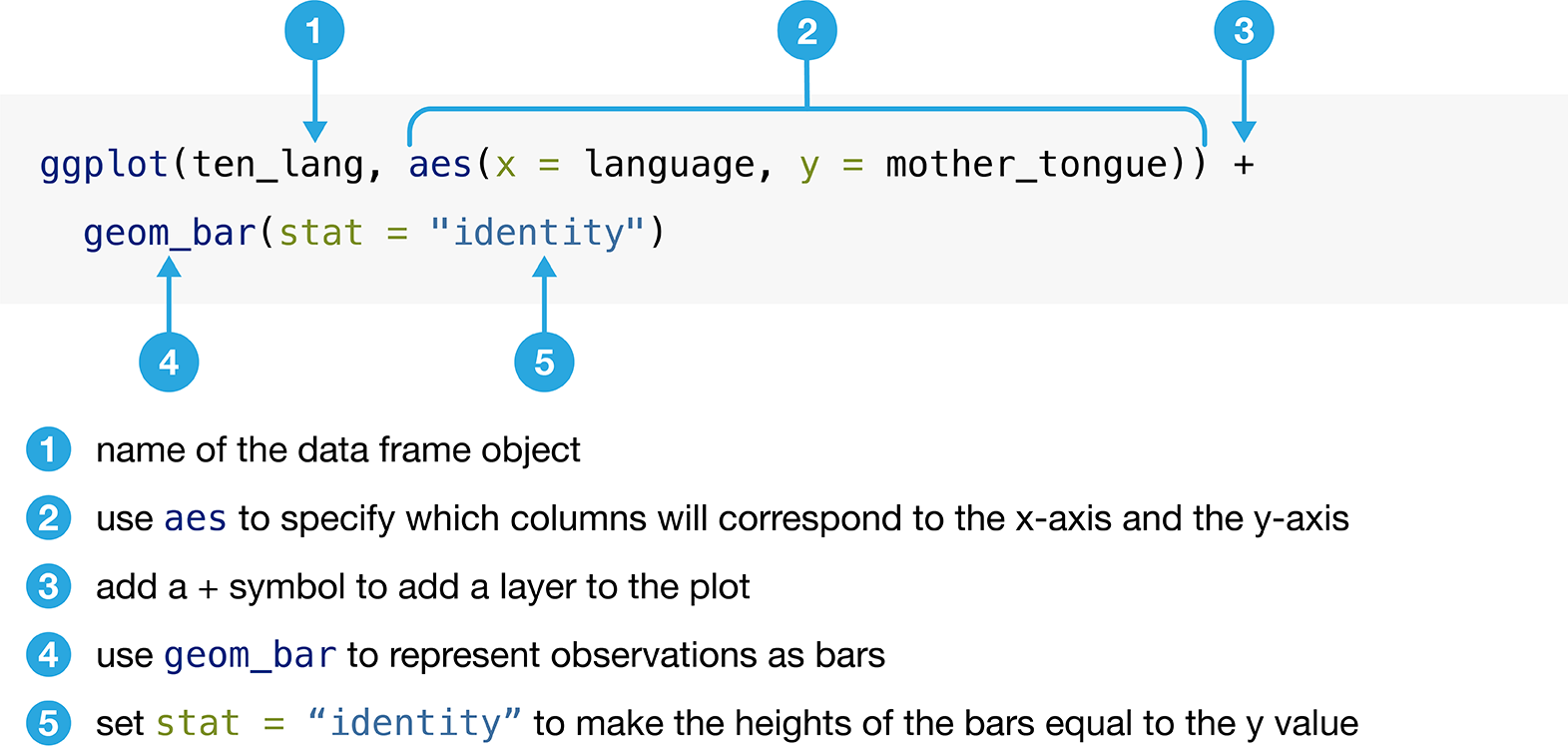
Figure 1.7: Creating a bar plot with the ggplot function.
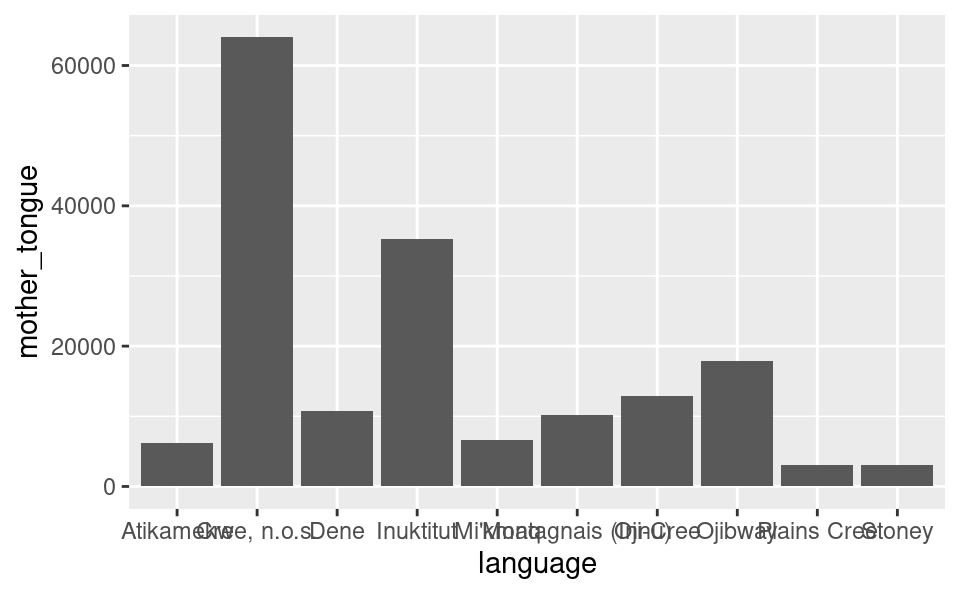
Figure 1.8: Bar plot of the ten Aboriginal languages most often reported by Canadian residents as their mother tongue. Note that this visualization is not done yet; there are still improvements to be made.
Note: The vast majority of the time, a single expression in R must be contained in a single line of code. However, there are a small number of situations in which you can have a single R expression span multiple lines. Above is one such case: here, R knows that a line cannot end with a
+symbol, and so it keeps reading the next line to figure out what the right-hand side of the+symbol should be. We could, of course, put all of the added layers on one line of code, but splitting them across multiple lines helps a lot with code readability.
1.10.2 Formatting ggplot objects
It is exciting that we can already visualize our data to help answer our
question, but we are not done yet! We can (and should) do more to improve the
interpretability of the data visualization that we created. For example, by
default, R uses the column names as the axis labels. Usually these
column names do not have enough information about the variable in the column.
We really should replace this default with a more informative label. For the
example above, R uses the column name mother_tongue as the label for the
y axis, but most people will not know what that is. And even if they did, they
will not know how we measured this variable, or the group of people on which the
measurements were taken. An axis label that reads “Mother Tongue (Number of
Canadian Residents)” would be much more informative.
Adding additional layers to our visualizations that we create in ggplot is
one common and easy way to improve and refine our data visualizations. New
layers are added to ggplot objects using the + symbol. For example, we can
use the xlab (short for x axis label) and ylab (short for y axis label) functions
to add layers where we specify meaningful
and informative labels for the x and y axes. Again, since we are specifying
words (e.g. "Mother Tongue (Number of Canadian Residents)") as arguments to
xlab and ylab, we surround them with double quotation marks. We can add many more
layers to format the plot further, and we will explore these in Chapter
4.
ggplot(ten_lang, aes(x = language, y = mother_tongue)) +
geom_bar(stat = "identity") +
xlab("Language") +
ylab("Mother Tongue (Number of Canadian Residents)")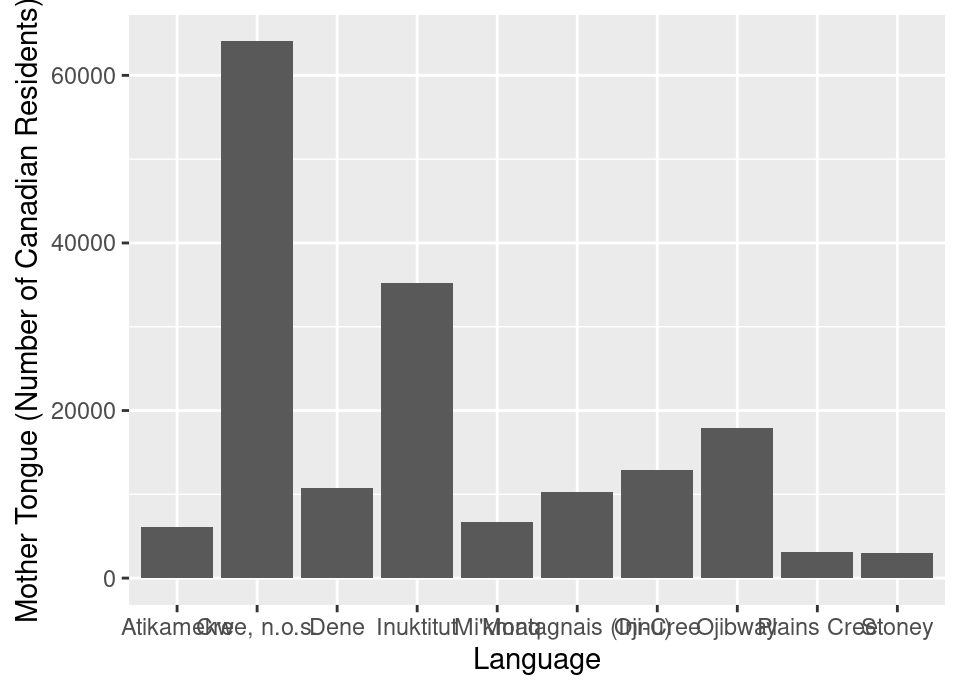
Figure 1.9: Bar plot of the ten Aboriginal languages most often reported by Canadian residents as their mother tongue with x and y labels. Note that this visualization is not done yet; there are still improvements to be made.
The result is shown in Figure 1.9. This is already quite an improvement! Let’s tackle the next major issue with the visualization in Figure 1.9: the overlapping x axis labels, which are currently making it difficult to read the different language names. One solution is to rotate the plot such that the bars are horizontal rather than vertical. To accomplish this, we will swap the x and y coordinate axes:
ggplot(ten_lang, aes(x = mother_tongue, y = language)) +
geom_bar(stat = "identity") +
xlab("Mother Tongue (Number of Canadian Residents)") +
ylab("Language")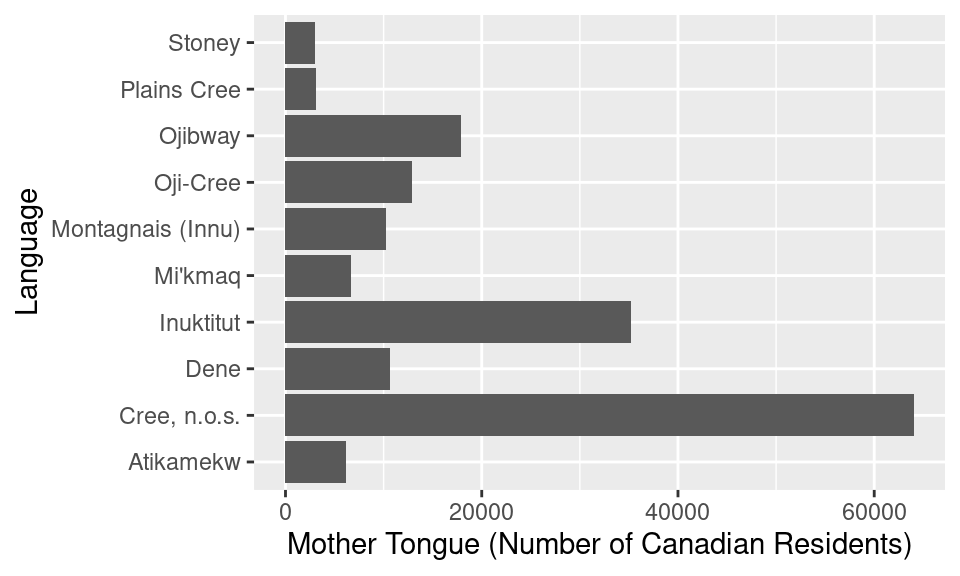
Figure 1.10: Horizontal bar plot of the ten Aboriginal languages most often reported by Canadian residents as their mother tongue. There are no more serious issues with this visualization, but it could be refined further.
Another big step forward, as shown in Figure 1.10! There
are no more serious issues with the visualization. Now comes time to refine
the visualization to make it even more well-suited to answering the question
we asked earlier in this chapter. For example, the visualization could be made more transparent by
organizing the bars according to the number of Canadian residents reporting
each language, rather than in alphabetical order. We can reorder the bars using
the reorder function, which orders a variable (here language) based on the
values of the second variable (mother_tongue).
ggplot(ten_lang, aes(x = mother_tongue,
y = reorder(language, mother_tongue))) +
geom_bar(stat = "identity") +
xlab("Mother Tongue (Number of Canadian Residents)") +
ylab("Language")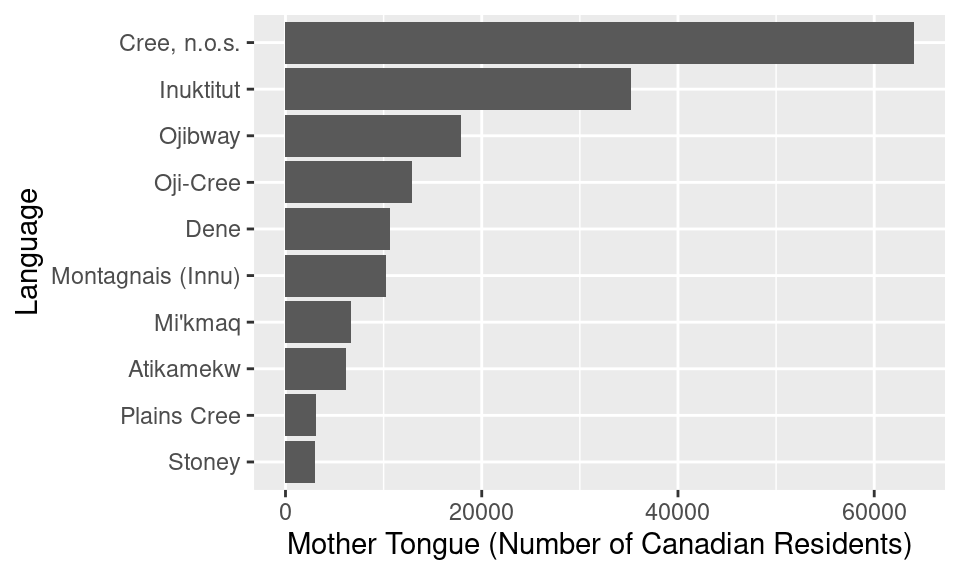
Figure 1.11: Bar plot of the ten Aboriginal languages most often reported by Canadian residents as their mother tongue with bars reordered.
Figure 1.11 provides a very clear and well-organized answer to our original question; we can see what the ten most often reported Aboriginal languages were, according to the 2016 Canadian census, and how many people speak each of them. For instance, we can see that the Aboriginal language most often reported was Cree n.o.s. with over 60,000 Canadian residents reporting it as their mother tongue.
Note: “n.o.s.” means “not otherwise specified”, so Cree n.o.s. refers to individuals who reported Cree as their mother tongue. In this data set, the Cree languages include the following categories: Cree n.o.s., Swampy Cree, Plains Cree, Woods Cree, and a ‘Cree not included elsewhere’ category (which includes Moose Cree, Northern East Cree and Southern East Cree) (Statistics Canada 2016b).
1.10.3 Putting it all together
In the block of code below, we put everything from this chapter together, with a few
modifications. In particular, we have actually skipped the
select step that we did above; since you specify the variable names to plot
in the ggplot function, you don’t actually need to select the columns in advance
when creating a visualization. We have also provided comments next to
many of the lines of code below using the
hash symbol #. When R sees a # sign, it
will ignore all of the text that
comes after the symbol on that line. So you can use comments to explain lines
of code for others, and perhaps more importantly, your future self!
It’s good practice to get in the habit of
commenting your code to improve its readability.
This exercise demonstrates the power of R. In relatively few lines of code, we
performed an entire data science workflow with a highly effective data
visualization! We asked a question, loaded the data into R, wrangled the data
(using filter, arrange and slice) and created a data visualization to
help answer our question. In this chapter, you got a quick taste of the data
science workflow; continue on with the next few chapters to learn each of
these steps in much more detail!
library(tidyverse)
# load the data set
can_lang <- read_csv("data/can_lang.csv")
# obtain the 10 most common Aboriginal languages
aboriginal_lang <- filter(can_lang, category == "Aboriginal languages")
arranged_lang <- arrange(aboriginal_lang, by = desc(mother_tongue))
ten_lang <- slice(arranged_lang, 1:10)
# create the visualization
ggplot(ten_lang, aes(x = mother_tongue,
y = reorder(language, mother_tongue))) +
geom_bar(stat = "identity") +
xlab("Mother Tongue (Number of Canadian Residents)") +
ylab("Language")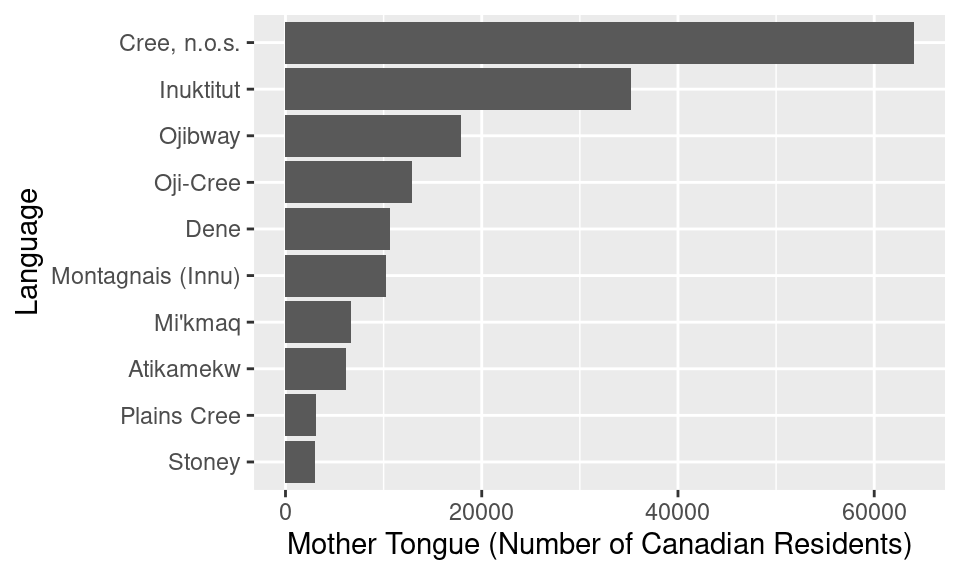
Figure 1.12: Putting it all together: bar plot of the ten Aboriginal languages most often reported by Canadian residents as their mother tongue.
1.11 Accessing documentation
There are many R functions in the tidyverse package (and beyond!), and
nobody can be expected to remember what every one of them does
or all of the arguments we have to give them. Fortunately, R provides
the ? symbol, which
provides an easy way to pull up the documentation for
most functions quickly. To use the ? symbol to access documentation, you
just put the name of the function you are curious about after the ? symbol.
For example, if you had forgotten what the filter function
did or exactly what arguments to pass in, you could run the following
code:
?filterFigure 1.13 shows the documentation that will pop up,
including a high-level description of the function, its arguments,
a description of each, and more. Note that you may find some of the
text in the documentation a bit too technical right now
(for example, what is dbplyr, and what is a lazy data frame?).
Fear not: as you work through this book, many of these terms will be introduced
to you, and slowly but surely you will become more adept at understanding and navigating
documentation like that shown in Figure 1.13. But do keep in mind that the documentation
is not written to teach you about a function; it is just there as a reference to remind
you about the different arguments and usage of functions that you have already learned about elsewhere.

Figure 1.13: The documentation for the filter function, including a high-level description, a list of arguments and their meanings, and more.
1.12 Exercises
Practice exercises for the material covered in this chapter can be found in the accompanying worksheets repository in the “R and the tidyverse” row. You can launch an interactive version of the worksheet in your browser by clicking the “launch binder” button. You can also preview a non-interactive version of the worksheet by clicking “view worksheet.” If you instead decide to download the worksheet and run it on your own machine, make sure to follow the instructions for computer setup found in Chapter 13. This will ensure that the automated feedback and guidance that the worksheets provide will function as intended.
 Data Science
Data Science